Evaluating and Enhancing Approaches for Improving the Robustness and Calibration of Neural Networks
Introduction
Electrocardiograms (ECGs) are widely used to detect heart diseases, monitor patients’ cardiovascular health over time, and support clinical decision making. However, automatic ECG interpretation remains challenging due to artifacts, differences in recording devices, and large variability between patients. Recent advances in deep learning offer the possibility to train general models that learn useful patterns from very large collections of ECGs and can later be adapted to many different diagnostic tasks. The goal of this project was to develop a general-purpose foundation model for ECGs that is robust, flexible, and transferable across many datasets and clinical scenarios. Instead of training a separate model for each task, a foundation model learns broad representations from large and diverse data sources. Training such a model requires processing millions of ECG segments and optimizing large neural networks. These requirements motivated the use of the High-Performance Cluster, which was essential to make this project feasible within a reasonable time frame due to its distributed training capabilities.
Methods
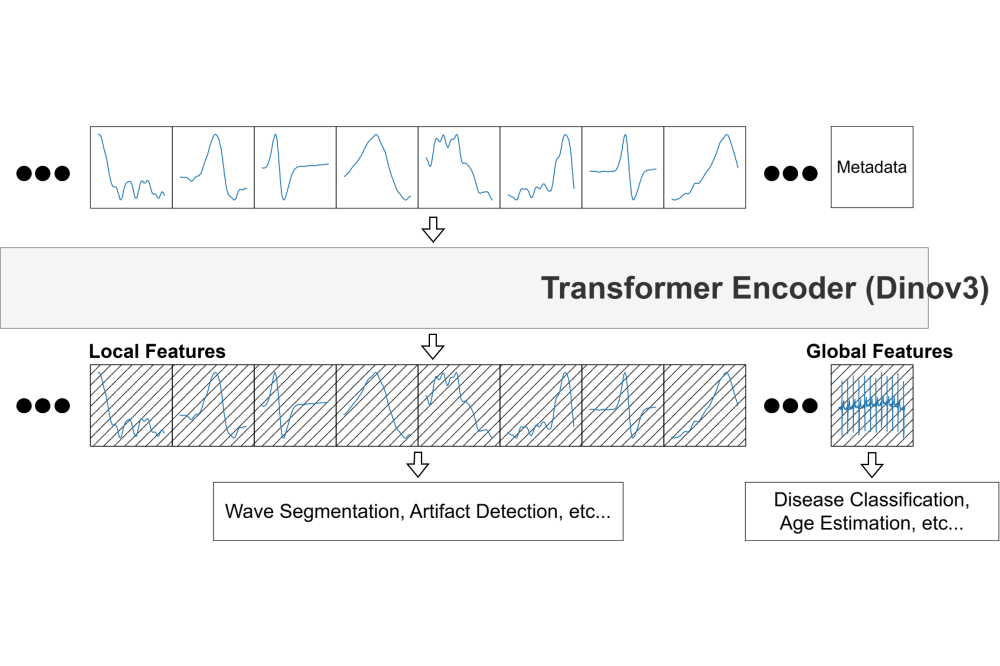
We adapted a modern self-supervised learning objective Dinov3, originally developed for images, to ECGs with an arbitrary number of leads. We opted for a Vision Transformer for our model architecture. The Dinov3 objective enabled our model to learn meaningful global and dense signal representations without relying on manual labels during training. This allowed the model to benefit from large datasets that would otherwise be too costly to annotate. To ensure robustness, we explicitly trained the model to handle common disturbances that appear in real-world recordings. These include electrode motion artifacts, baseline drifts, and muscle interference. We achieved this by augmenting the training data with realistic noise patterns taken from the MIT-BIH noise stress test database. As a result, the model learned to focus on clinically relevant signal features rather than being distracted by noise. Our training dataset combined several large and diverse collections of ECGs originating from different hospitals, devices, and recording protocols: The Harvard Emory ECG Database, the CODE database, and the Incentia11k database. Importantly, the data included recordings with varying numbers of leads, which allowed the model to generalize across different sensor configurations. Training the model required distributing both the data and its parameters across multiple CUDA-enabled GPUs. We used the PyTorch library and a fully sharded data parallel approach to efficiently handle memory usage and computation.
Results
We evaluated the learned representations on multiple downstream tasks to assess their quality. First, we tested global multilabel classification performance by linearly probing the frozen model representations using the PTB-XL dataset and achieved an Area Under the Receiver Operating Characteristic Curve (AUROC) of 0.87 on the test set. Second, we evaluated fine-grained classification performance at the level of individual heartbeats using the MIT-BIH Arrhythmia database and achieved an AUROC of 0.80.
Discussion
The results show that self-supervised foundation models can effectively learn general representations from large collections of Electrocardiograms. By explicitly incorporating realistic artifacts during training and using diverse datasets, we achieved a model that generalizes well across tasks, recording devices, and signal configurations. This robustness is particularly important for real-world clinical applications, where ideal recording conditions cannot be guaranteed. In future work, we plan to extend the model to handle much longer recordings by increasing its temporal context. Additionally, we aim to incorporate multiple data modalities, such as combining Electrocardiograms with Echocardiograms or clinical text. These extensions will further increase computational demands, reinforcing the importance of the High-Performance Cluster for this line of research.