MD-Simulations to Study the Effect of Electric Fields on the SARS-CoV-2 Postfusion Structure
Introduction
The interaction between the Spike (S) glycoprotein and the ACE2 receptor on the host cell constitutes the pivotal initial phase of virus entry. In its prefusion state, the S protein, similar to other class 1 viral fusion proteins, displays metastability, a characteristic that significantly influences its optimization and regulation of functions. The prefusion S protein contains both S1 and S2 subunits, which remain noncovalently bonded until viral fusion is initiated. The subsequent dissociation of S1 sets off a sequence of refolding events in the metastable prefusion S2, facilitating the fusogenic transition to a stable postfusion structure. In a previous projects, employing molecular dynamics (MD) simulations, we demonstrated that low to moderate electric fields with intensities as low as 105 V/m can affect the Receptor
Binding Domain in wild type (WT), Alpha, Beta, Gamma, Delta and Omicron variants. This modification of the S protein enables it to overcome a non-thermal energy barrier, thereby shifting it to a conformational state between the prefusion and postfusion states. In the current project, we assess the impact of external electric fields (EFs) on the conformational stability of the postfusion S2 non-functional form through MD simulations, considering its dynamic structural changes. Interestingly, we observed no significant structural alterations, even when applying high-intensity EFs (108 V/m), which exceeds the threshold for significant conformational changes reported in many globular proteins.
Methods
The initial proteins conformations were derived from available Protein Data Bank (PDB) structures. The simulations were performed using the GROMACS package (version 2019.4) with CHARMM-36 force field parameters. The resulting trajectory files were analyzed using GROMACS tools and the MDAnalysis Python library. The Adaptive Poisson-Boltzmann Solver (APBS) algorithm was used to calculate all potential maps on selected frames from the MD trajectories. Molecular docking calculations for protein-receptor interactions were carried out using the pyDock web server. Visualization of the MD trajectories and molecular representations was accomplished using the VMD software.
Results
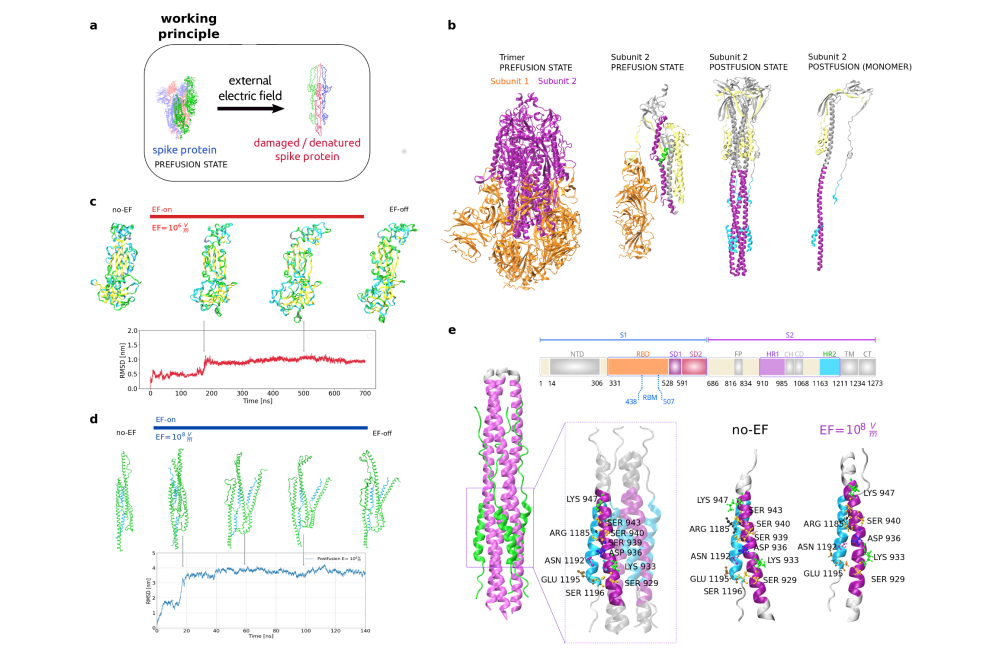
The impact of external electric fields (EFs) on the overall structure of the S1 and S2 subunits of the S protein was investigated through molecular dynamics simulations. The results of our simulations shown significant differences in the global structural changes in the S1 and S2 Subunits of the Spike Protein under Electric Fields. To examine the prefusion S1 subunit, we applied low field intensities ranging from 104 to 107 V/m, building upon our previous findings. We observed S1 protein elongation in the trajectories due to the alignment of permanent local dipoles and the displacement of charges parallel to the electric field. This also led to a loss of tertiary protein structures within a few nanoseconds. The conformational shifts of the prefusion S1 subunit protein under the influence of the electric field are evident from the time-dependent changes in the root-mean-square displacements (RMSD) of the protein backbone relative to the initial structures. The transition from the initial conformation to a new stable structure occurs within the first 200 nanoseconds. Simulations for the stable postfusion S2 subunit were conducted with an EF intensity of 108 V/m, slightly below the damage threshold for globular proteins. We found that the postfusion state of the S protein, under a field strength of 108 V/m, exhibits significantly greater stability compared to the metastable prefusion state. The overall three-dimensional structure underwent changes during the initial 20 nanoseconds of the dynamics while maintaining the interaction between regions in each protomer. We can observe in the RMSD that this conformation remains stable throughout the entire simulation without significant additional changes. Furthermore, our simulations demonstrate that the postfusion conformation does not experience irreversible damage to its secondary structure even when subjected to electric fields as high as 108 V/m.
Discussion
In this project, our simulations focused on the prefusion and postfusion conformations to characterize the differences in the stability and vulnerability of both structures. We found that the receptor binding domain of the prefusion conformation of the spike protein undergoes significant changes in the tertiary as well as the secondary structure under the application of EFs. We also demonstrate that, in contrast, the S2 subunit of the postfusion state of the spike protein remains structurally intact even when exposed to electric field intensities four orders of magnitude higher. The postfusion conformation displays changes only in the tertiary structure, which can be attributed to its slender shape that contributes to its flexibility. These findings offer a strong scientific foundation to affirm the correlation between the prefusion-state metastability of the SARS-CoV-2 spike protein and its vulnerability to electric field damage. Following the virus attachment to the ACE2 receptor, the spike protein assumes a stable and resilient postfusion conformation, exhibiting resistance to electric fields (with a damage threshold exceeding 108 V/m), similar to that observed in most globular proteins.