Mapping the Genetics of Neuropsychological Traits to Molecular NETworks of Human Brain
Introduction
The complex architecture of neuropsychiatric disorders comprises of distinct neuropsychological traits. Previous studies have shown that these neuropsychological traits are modulated by diverse genetic factors. To further explore and untangle the association between such complex traits and the underlying genetics genome-wide association studies (GWAS) are performed. These studies require thorough quality check of the genetic data, imputation of missing data and in-depth downstream analysis. However, in the area of neuropsychology there is no standalone tool existing which takes raw genetic data (pre-quality check) and a quantitative neuropsychological trait that besides performing quality check, imputation and genome-wide association analysis also maps those genetic association at brain level.
Methods
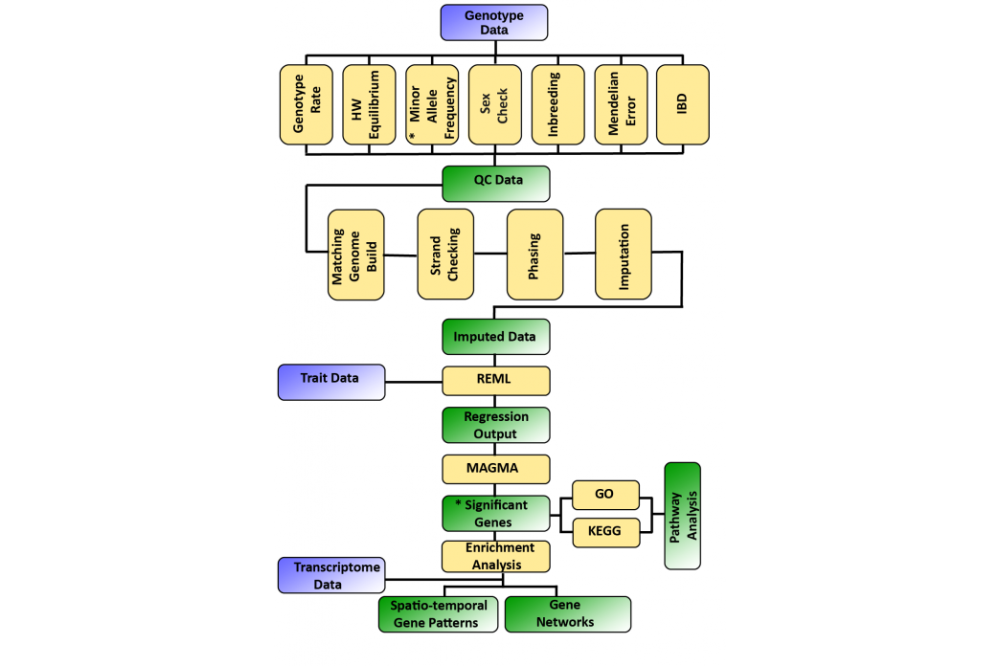
We develop a pipeline “MAGNET” that integrates the genetic data (SNP: Single nucleotide polymorphism), trait data (e.g. Intelligence Quotient (IQ) scores) and brain transcriptome data (sum of all Ribonucleic acid (RNA) molecules) on one platform. MAGNET is written in R-3.4.4, bash and python. The framework is currently being developed under Linux distribution red hat 4.1.2-55. We implemented it on an autism spectrum disorders (ASD) dataset using IQ data as the neuropsychological trait. The calculations are performed on “FUCHS-CSC” high performance cluster of the Goethe University Frankfurt. MAGNET workflow is presented in Figure 1 below.
Results
Based on the ASD data we identify the genetic variants which are associated with IQ and similarly their respective genes. We determine that the IQ related genes are related to gene networks which are down-regulated in early post-conception weeks and immediately after birth, but are up-regulated after 2 years of age implicated in frontal cortex and striatum.
Discussion
We provide a comprehensive pipeline which integrates state-of-the-art algorithms. It provides extensive analysis starting from a neuropsychological trait to its association with specific brain areas and time points within one shell. This can assist researchers to gather a meaningful interpretation of their trait of interest within a time span of four to five days.
Outlook
MAGNET is currently under development and will be available as a command line tool only. In the near future we will develop a graphical user interface which can provide a more user friendly environment.